3.5: Cre-Lox, Driver Lines, and Next Order Specificity
- Page ID
- 10653
What is the Cre-lox system and why do we need it?
In the previous chapters we have examined the use of recombinant molecular techniques that help neurobiologists to understand the role of specific neurons within a brain circuit in a complex behavior. However, we haven’t yet examined how to ensure delivery to specific neurons to ensure that the construct (i.e.ChR2, hM3DR or other constructs) is not inappropriately expressed in other cell types nor other neurons. Instead of relying only on the cells internal machinery being driven by cell specific gene promoters, the Cre-lox system is often used in neuroscience. This is a powerful method to selectively manipulate gene expression in specific subsets of tissues. There are 2 elements to make this system work effectively; loxP sequences that are created de novo and the presence of Cre-recombinase (an exogenous nuclease). Importantly neither the nuclease enzyme Cre recombinase nor loxP sequences are normally found in mammal systems, so their introduction has no functional consequences to the DNA nor to the neurons. Genetically engineered loxP sites are 34 bp DNA sequences that bind to the protein Cre-recombinase (and the nuclease always needs to bind to 2 loxP sites). Interestingly, as shown in Figure \(\PageIndex{1}\), when loxP sites are in the same orientation (5’ to 3’) and Cre-recombinase binds to both, this causes deletion of the DNA sequence between the loxP sites while leaving one loxP. This type of “floxing” reaction is known as excision. If the loxP sites are in the opposite orientation then the sequence between them is inverted or “flipped”.
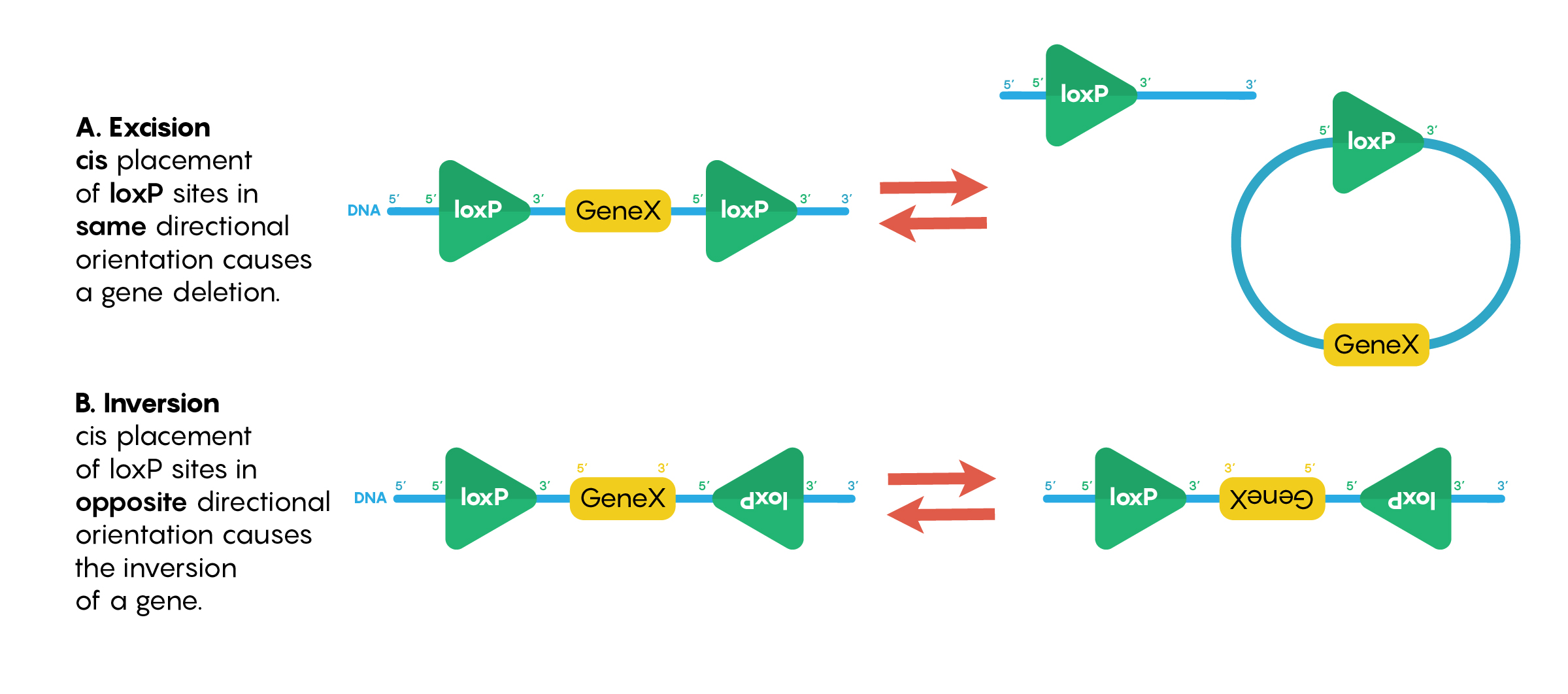
Figure \(\PageIndex{1}\). Different orientations of the loxP sites around the DNA sequence of interest will cause either excision (removal of the DNA) or inversion of the DNA sequence.
Cre-driver lines: A special type of transgenic mouse
As Cre-recombinase has no effect on DNA except for those that contain loxP sites (which are never found in mammalian DNA), generating transgenic Cre-recombinase mouse models was a logical step in examining specific expression of genetic tools (optogenetic, chemogenetic or other) in neurons. As different neurons have different cellular machinery to activate and express certain genes, Cre-recombinase was targeted to specific neuronal (or other brain cell types) with this in mind. As the Cre-recombinase expression was being “driven” by the machinery specific to a cell type, the term Cre-driver lines has become embedded within neurobiology. As shown in Figure \(\PageIndex{2}\) below, different types of neurons, whether inhibitory, excitatory or other have specific proteins and therefore genes that would allow for the specific expression of the exogenous Cre-recombinase. For example the TH::IRES-Cre-recombinase mouse only expresses Cre-recombinase within dopaminergic neurons as TH (Tyrosine Hydroxylase) is the synthetic enzyme that converts the amino acid tyrosine to dopamine in neurons. In the same vein, VGAT::IRES-Cre-Recombinase would be a transgenic mouse line that would be used to investigate inhibitory GABAergic neurons. A list of different genes/protein products is found in both Figure \(\PageIndex{2}\) and the following table.
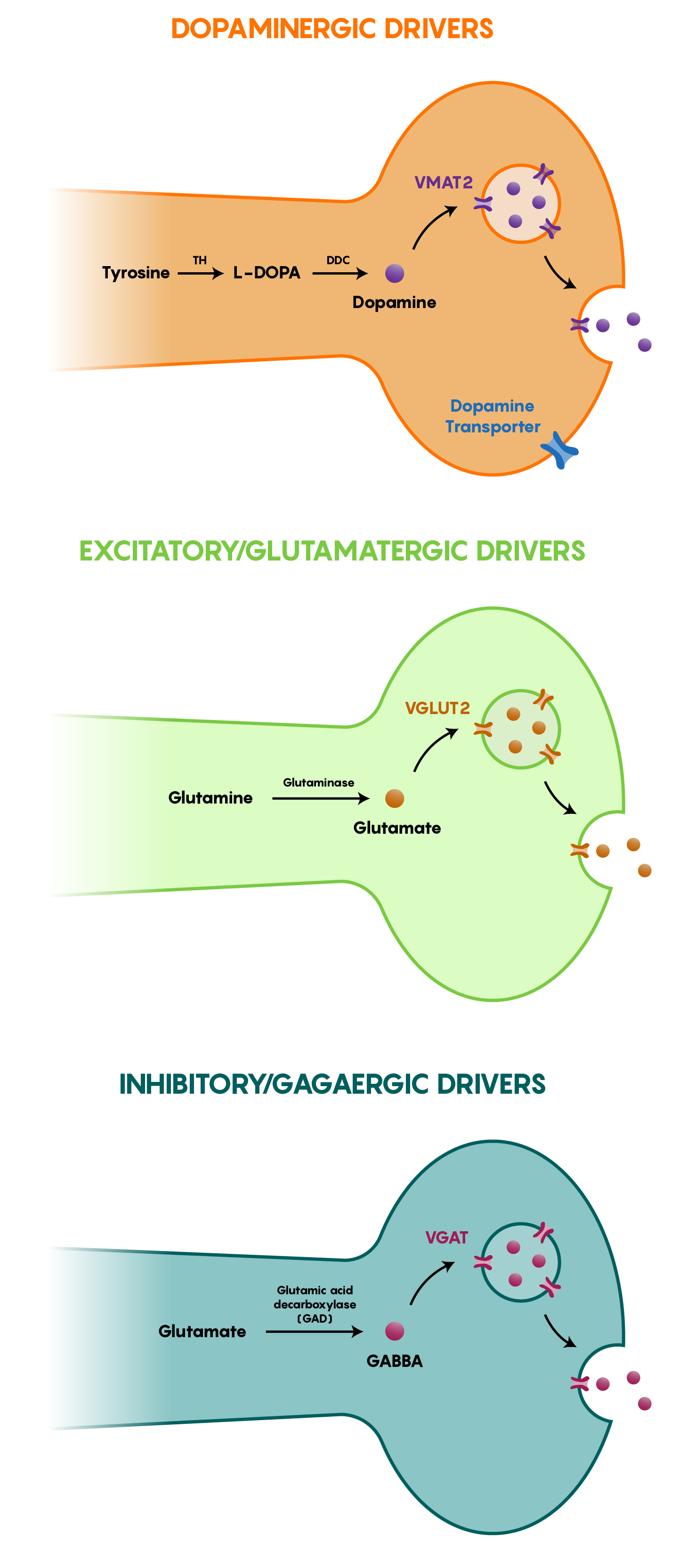
Figure \(\PageIndex{2}\). Differential expression of proteins in specific groups of neurons including dopaminergic, excitatory glutamatergic and inhibitory GABAergic types that allow for the directed expression of Cre-recombinase in specific subsets of neurons.
| Driver line | Specific cell types – Cre-Recombinase |
|---|---|
| GFAP-Cre | Astrocytes or neuronal stem cells |
| Nestin-Cre | Neuronal stem cells or precursor cells |
| CAG-Cre | General promoter: Ubiquitous expression in all cells of the brain |
| Synapsin I-Cre | All neurons (all types of neurons |
| EF1a-Cre | General promoter: Ubiquitous expression in all cells of the brain |